For å lese pluss-artikler må du være abonnent
Et abonnement gir tilgang til alt innhold og vi har følgende tilbud
Insight
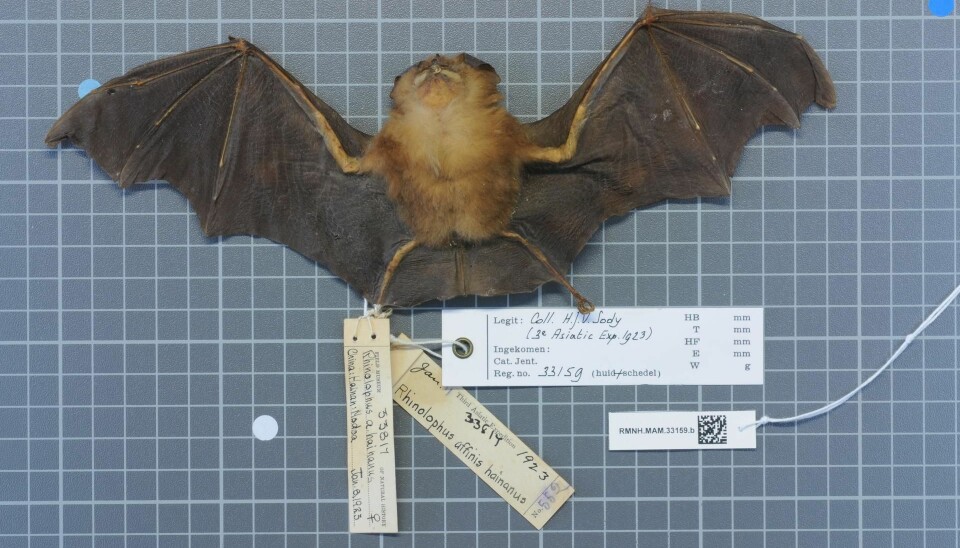

The virus most similar to SARS-CoV-2 was discovered in a mineshaft in 2013 after several contracted a mysterious lung disease. Wuhan's Institute of Virology first claimed that this virus was not sequenced until 2020, but now head researcher Shi Zhengli offers another explanation.
On February 3rd, 2020, researchers at the Wuhan Institute of Virology published their first article on the SARS-CoV-2 virus.
In the article “A pneumonia outbreak associated with a new coronavirus of probable bat origin”, the Wuhan researchers Zhou et al. compare the SARS-CoV-2 virus with other known viruses. Zhou et al also published the sequence of a hitherto completely unknown virus, called RaTG13, which is 96.2 percent identical to SARS-CoV-2 virus.
The article has already been cited 3261 times, and is undoubtedly one of the most influential scientific publications in 2020.
Among the articles that builds upon the research published by the Wuhan Institute of Virology, and more specifically the RaTG13 sequence, is the article “The proximal origin of SARS-CoV-2” by Kristian Andersen et al. This article is one of the most influential scientific articles arguing for a natural origin for the novel coronavirus. Andersen’s article is cited over 400 times so far in 2020.
However, some researchers are in doubt as to whether the raw data this research builds upon is credible. At the heart of the issue is the RaTG13 virus, and whether the researchers in Wuhan have provided sufficient and correct information about the sequencing of the virus.
Shortly after publication, questions about the hitherto unknown virus began to surface. On May 19th, the Indian researchers Monali C. Rahalkar and Rahul A. Bahulikar published an article claiming that the RaTG13 virus is identical to a virus sample named BtCoV/4991, which was uploaded by the Wuhan researchers in 2016.
The BtCoV/4991 virus sample consists only of 370 fragments out of the more than 30,000 positions that completes a SARS-coronavirus genome. The 370 fragments in the BtCov/4991 virus sample make a 98.9 percent match with the certain positions in the genome of the SARS-CoV-2 virus, deviating only by four out of 370 positions.
Rahalkar and Bahulikar criticize the researchers in Wuhan for not making it clear in their article that the RaTG13 virus is taken from BtCoV/4991 sample, and for failing to mention that this sample was collected from an abandoned mineshaft. This is of particular interest since six workers from this mineshaft were hospitalized after having contracted an unknown disease that killed three of them.
Admittedly, chief researcher Shi Zhengli, at the Wuhan Institute of Virology, has mentioned this incident earlier. In an interview with The Scientific American, she explains how her team of researchers were called to investigate the mine after the incident with the sick workers.
According to Shi’s explanation, the investigation of the events concluded that the disease was caused by fungi.
However, a master’s thesis written in 2013 by one of the doctors who assisted in the treatment of the miners, casts doubt on Shi Zhengli’s explanation.
The thesis describes a course of the disease very similar to what patients who are now diagnosed with Covid-19 experience. The patients were tested for several known diseases, but all the tests came back negative. The thesis further discredits the notion that the patients suffered from fungal infection, stating that one «patient did not receive any anti-fungal medicine for treatment, yet still recovered. This suggested that the possibility of the illness being triggered by fungal infection is slim».
Furthermore, the thesis states that the doctors that worked to treat the sick miners «worked with Dr. Zhong Nan Shan and did some sampling». Dr. Zhong Nan Shan is one of China's foremost experts on SARS-coronaviruses, and led his country's response to the 2003 SARS epidemic.
The Wuhan Institute of Virology is also mentioned in the paper, as one «patient tested positive for Serum IgM by the WuHan Institute of Virology», which «suggested the existence of virus infection». The master thesis concludes that the unknown virus leading to severe pneumonia could be either «the SARS-like-CoV from the Chinese rufous horseshoe bat or Bats kind SARS-like CoV».
The ambiguity surrounding the sick miners, the disease they suffered from and the virus samples taken from the mine shaft and shipped to Wuhan in 2013, has therefore triggered a flood of questions for Shi Zhengli and her research team at the Wuhan Institute of Virology.
And in an interview with ScienceMag, Zhengli answers multiple questions relating to the origin of these viruses. Here she confirms that RaTG13 and BtCoV/4991 refer to the same virus. She also explains that the name BtCoV/4991 refers to the sample the virus is taken from, while RaTG13 is the name of the virus itself.
When asked about when Shi and her colleagues sequenced the entire RaTG13 virus, Shi Zhengli answers that this was done in 2018, and that the only virus sample that contained the virus was used up after sequencing. According to this explanation, the laboratory in Wuhan has therefore not stored the virus since 2018:
In 2018, as the NGS sequencing technology and capability in our lab was improved, we did further sequencing of the virus using our remaining samples, and obtained the full-length genome sequence of RaTG13 except the 15 nucleotides at the 5’ end. As the sample was used many times for the purpose of viral nucleic acid extraction, there was no more sample after we finished genome sequencing, and we did not do virus isolation and other studies on it. Among all the bat samples we collected, the RaTG13 virus was detected in only one single sample. In 2020, we compared the sequence of SARS-CoV-2 and our unpublished bat coronavirus sequences and found it shared a 96.2% identity with RaTG13. RaTG13 has never been isolated or cultured.
But this explanation differs from what Wuhan’s Institute of Virology wrote in the article they published in Nature on February 2nd. Here they instead write the following about when the virus was sequenced:
We then found that a short region of RNA-dependent RNA polymerase (RdRp) from a bat coronavirus (BatCoV RaTG13) —which was previously detected in Rhinolophus affinis from Yunnan province — showed high sequence identity to 2019-nCoV. We carried out full-length sequencing on this RNA sample.
This makes Alina Chan, a postdoctoral fellow at the Broad Institute of MIT and Harvard, where she works with molecular biology and gene therapy, react:
“Many readers of the Nature article, including myself, interpreted the relevant text to mean that the RaTG13 full-length genome sequencing was only performed after the RdRp match between SARS-CoV-2 and RaTG13. From interviews of Peter Daszak, a close collaborator of Shi Zhengli, I also had the impression that the RaTG13 sample had not been full-genome sequenced until after the COVID-19 outbreak. However, in the Science Q&A, Shi informed us that RaTG13's full genome had been sequenced in 2018, and that this process had depleted the sample entirely”, she explains to Minerva.
Alina Chan further adds that the methods section of the Nature article looks strange, if the Wuhan Institute of Virology had already completed sequencing of RaTG13 in 2018.
In this section of the paper, the scientists at the Wuhan Institute of Virology, Zhou et al. write that in order to sequence the SARS-CoV-2 virus, they «aligned reads to a local database».
Then «by de novo assembly and targeted PCR», the Wuhan Scientist «obtained a 29,891-base-pair CoV genome that shared 79.6% sequence identity to SARS-CoV BJ01 (GenBank accession number AY278488.2).»
"Given that they already had the full genome sequence of RaTG13 back in 2018, wouldn't they have immediately found the 96.2% genome identity match upon querying their internal database of virus sequences for matches to the de novo assembled SARS-CoV-2 genome?" she ponders.
“Surprisingly, they wrote that they found a 79.6% genome match to SARS-CoV BJ01, and a close match to a short region of the RdRp of RaTG13 ––– instead of writing straight away that they found a whopping 96.2% genome match to RaTG13", Chan elaborates.
Another unresolved question raised by this is which database the RaTG13 sequence was uploaded to in 2018, and if other unpublished virus genomes are stored on this local database.
Gunnveig Grødeland is a researcher at the Department of Immunology and Transfusion Medicine at the University of Oslo. Like Chan, she sees a contradiction in Shi Zhengli’s statements about when the virus was sequenced.

“This is a clear contradiction that should be investigated”, she says.
On why the timing of the sequencing of RaTG13 virus matters, Grødeland responds:
“For the quality of the sequencing, it does not, but for the discussion about the origin, this is relevant. One of the central principles in research is transparency about the information that is available, including what work has been done and how this was done.”
“This may have a natural explanation, but it could also be that it does not. That is why we need more transparency on the raw data”, she elaborates.
Asked about whether we should be concerned about the quality of the raw data provided by the Wuhan Institute of Virology on the RaTG13 sequence, Grødeland says:
“One of the things that should definitely be avoided is conspiracy theories, but asking relevant critical questions is what research is all about. So far, the origin of the SARS-CoV-2 virus is unknown, and something should be investigated. It should at least be in Wuhan’s Institute of Virology's self-interest to practice full transparency about its raw data, the work that has been done at the institute, and when this was conducted.”
On August 6th, Minerva presented the contradiction between, one the one hand, Shi Zhengli’s statements to ScienceMag, and on the other, and what was written in the February article in Nature, for the editorial staff of Nature. The editorial staff was asked whether the journal had made any attempt to clarify or rectify the content of the article given that Shi Zhengli has now given a different explanation as to when and why RaTG13 was sequenced.
On the same day, Minerva received a reply from Nature that the inquiry would be taken up with the journal’s editor, and that it was not possible to give further comments at that time. After some time a spokesperson for the journal responded on August 18th:
“We look into any comments or concerns raised about any Nature paper in detail, including those regarding methodological details. In general, our editors will assess comments or concerns that are raised with us in the first instance, consulting the authors and, where appropriate, seeking advice from peer reviewers and other external experts. We are currently considering comments that have been raised with us relating to this paper, and cannot comment further at this time.”